Theory pages.
Over the last few days I've been trying to teach myself enough genetics to reconstruct Carrion-Vazquez's poly-I27 synthesis procedure. I'm not quite there yet, but I feel like I've made enough progress that it's worth posting my notes somewhere public in case they are useful to others.
Overview
We buy our poly-I27 from AthenaES, who market it as I27O™. Perusing their technical brief, makes it clear that I2O7™ corresponds to Carrion-Vazquez's I27RS₈. In Carrion-Vazquez' original paper they describe the synthesis of both I27RS₈ and a variant I27GLG₁₂. Their I27RS₈ procedure is:
- Human cardiac muscle used to generate a cDNA library (Rief 1997)
- cDNA library amplified with PCR
- 5' primer contained a BamHI restriction site that permitted in-frame cloning of the monomer into the expression vector pQE30.
- The 3' primer contained a BglII restriction site, two Cys codons located 3' to the BglII site and in-frame with the I27 domain, and two in-frame stop codons.
- The PCR product was cloned into pUC19 linearized with BamHI and SmaI.
- The 8-domain synthetic gene was constructed by iterative cloning of monomer into monomer, dimer into dimer, and tetramer into tetramer.
- The final construct contained eight direct repeats of the I27 domain, an amino-terminal His tag for purification, and two carboxyl-terminal Cys codons used for covalent attachment to the gold-covered coverslips.
They also give the full-length sequence of I27RS₈:
Met-Arg-Gly-Ser-(His)₆-Gly-Ser-(I27-Arg-Ser)₇-I27-...-Cys-Cys
They point out the Arg-Ser (RS) amino acid sequence is the BglII/BamHI hybrid site, which makes sense.
Back on the Athena site, they have a page describing their procedure (they reference the Carrion-Vazquez paper). They claim to use the restriction enzyme KpnI in addition to BamHI, BglII, and SmaI.
Carrion-Vazquez points to the following references:
- Kempe et al. 1985 (CV16), the source of the multi-step cloning technique.
- Rief et al. (CV10), for I27 subcloning.
Rief
In their note 11, Rief et al. explain their synthesis procedure:
- λ cDNA library
- Titin fragments of interest were amplified by PCR
- cloned into pET 9d
- NH₂-terminal domain boundaries were as in Politou 1996.
- The clones were fused with an NH₂-terminal His₆ tag and a COOH-terminal Cys₂ tag for immobilization on solid surfaces.
which doesn't help me very much.
Kemp
The Kempe article is more informative, focusing entirely on the synthesis procedure (albiet for a different gene). Their figure 2 outlines the general approach, and used the following restriction enzymes: PstI, BamHI, PstI, and BglII. I'll walk through their procedure in detail below.
Genetic code
Wikipedia has a good page on the genetic code for converting between DNA/mRNA codons and amino acids. I've written up a little Python script, mRNAcode.py, to automate the conversion of various sequences, which helped me while I was writing this post. I'm sure there are tons of similar programs out there, so don't feel pressured to use mine ;).
Restriction enzymes
We'll use the following restriction enzymes:
5' G|GATC C 3'
3' C CTAG|G 5'
BglI (N is any nucleotide)
5' GCCN NNN|NGGC 3'
3' CGGN|NNN NCCG 5'
5' A|GATC T 3'
3' T CTAG|A 5'
5' A|AGCT T 3'
3' T TCGA|A 5'
5' G GTAC|C 3'
3' C|CATG G 5'
5' C TGCA|G 3'
3' G|ACGT C 5'
5' CCC|GGG 3'
3' GGG|CCC 5'
Details
Here's my attempt to reconstruct the details of the polymer-cloning reactions, where they splice several copies of I27 into the expression plasmid.
Kempe procedure
Inserted their poly-SP into pHK414 (I haven't been able to find any online sources for pHK414. Kempe cites R.J. Watson et al. Expression of Herpes simplex virus type 1 and type 2 glyco-protein D genes using the Escherichia coli lac promoter. Y. Becker (Ed.), Recombinant DNA Research and Viruses. Nijhoff, The Hague, 1985, pp. 327-352.)
Synthetic SP
HindIII. ,BamHI_.
| | Met Arg Pro Lys Pro Gln Gln Phe Phe Gly Leu Met |
5’ GA AGC TTC ATG CGT CCG AAG CCG CAG CAG TTC TTC GGT CTC ATG GAT CCG
CT TCG AAG TAC GCA GGC TTC GGC GTC GTC AAG AAG CCA GAG TAC CTA GGC 5’
pHK414
_______Linker_sequence______
/ \
HindIII BamHI
,PstI. BglII.| |,SmaI. |
CTGCAG...AGATCTAAGCTTCCCGGGGATCCAAGATCC
GACGTC...TCTAGATTCGAAGGGCCCCTAGGTTCTAGG
. .
.......................................
Synthesizing pSP4-1
pHK414 + HindIII + BamHI
They cut a hole in the plasmid…
HindIII BamHI.
(PstI) BglII,| |
CTGCAG...AGATCTA GATCCAAGATCC
GACGTC...TCTAGATTCGA GTTCTAGG
. .
.......................................
SP + HindIII + BamHI
… and cut matching snips off their SP gene.
HindIII. ,BamHI_.
| | Met Arg Pro Lys Pro Gln Gln Phe Phe Gly Leu Met |
AGC TTC ATG CGT CCG AAG CCG CAG CAG TTC TTC GGT CTC ATG
AG TAC GCA GGC TTC GGC GTC GTC AAG AAG CCA GAG TAC CTA G
pSP4-1
Mixing the snips together gives the plasmid with a single SP.
HindIII BamHI.
,PstI. BglII.| | MetArgProLysProGlnGlnPhePheGlyLeuMet |
CTGCAG...AGATCTAAGCTTCATGCGTCCGAAGCCGCAGCAGTTCTTCGGTCTCATGGATCCAAGATCC
GACGTC...TCTAGATTCGAAGTACGCAGGCTTCGGCGTCGTCAAGAAGCCAGAGTACCTAGGTTCTAGG
. .
......................................................................
Using -SP- to abbreviate the HindIII→Met→Met portion (less the
terminal G, which is part of the BamHI match sequence).
,PstI. BglII. BamHI.
CTGCAG...AGATCT-SP-GGATCC
GACGTC...TCTAGA-SP-CCTAGG
. .
.........................
Synthesizing pSP4-2
The single-SP plasmid, pSP4-1, is split in two parallel reactions.
PstI + BamHI
G...AGATCT-SP-G
ACGTC...TCTAGA-SP-CCTAG
PstI + BglII
CTGCA GATCT-SP-GGATCC
G A-SP-CCTAGG
. .
.........................
pSP4-2
Then the SP-containing fragments (shown above) are isolated and mixed together to form pSP4-2.
,PstI. BglII. other. BamHI.
CTGCAG...AGATCT-SP-GGATCT-SP-GGATCC
GACGTC...TCTAGA-SP-CCTAGA-SP-CCTAGG
. .
...................................
where the "other" sequence is the result of the BamHI/BglII splice.
Expanding the -SP- abbreviation around the SP joint:
....SP,other_.HindIII. SP.....
Leu Met Asp Leu Ser Phe Met Arg
CTC ATG GAT CTA AGC TTC ATG CGT
AGA CGT TCG AGC CTA GGA CGT ATG
So the resulting poly-SP will have Asp-Leu-Ser-Phe linking amino acids.
By repeating the PstI + BamHI / PstI + BglII split-and-join, you can synthesize plasmids with any number of SP repeats.
I27RS₈ procedure
Like Kempe, Carrion-Vazquez et al. flank the I27 gene with BglII and BamHI, but they reverse the order. Here's the output of their PCR:
BamHI-I27-BglII-Cys-Cys-STOP-STOP
From the PDB entry for I27 (1TIT), the amino acid sequence is:
,leader_.
MHHHHHHSSLIEVEKPLYGVEVFVGETAHFEIELSEPDVHGQWKLKGQPLTASPDCEIIEDGKKHILI
LHNCQLGMTGEVSFQAANAKSAANLKVKEL
To translate this into cDNA, I've scanned thorough the sequence of NM_003319.4, and found a close match from nucleotides 15991 through 16248.
15982 CTAATAAAAG TGGAAAAGCC TCTGTACGGA GTAGAGGTGT TTGTTGGTGA
16032 AACAGCCCAC TTTGAAATTG AACTTTCTGA ACCTGATGTT CACGGCCAGT
16082 GGAAGCTGAA AGGACAGCCT TTGACAGCTT CCCCTGACTG TGAAATCATT
16132 GAGGATGGAA AGAAGCATAT TCTGATCCTT CATAACTGTC AGCTGGGTAT
16182 GACAGGAGAG GTTTCCTTCC AGGCTGCTAA TGCCAAATCT GCAGCCAATC
16232 TGAAAGTGAA AGAATTG
This cDNA match generates an amino acid starting with LIKVEK instead of the expected LIEVEK, but the LIKVEK version matches amino acids 12677-12765 in Q8WZ42 (canonical titin), and there is a natural variant listed for 12679 K→E.
Interestingly, this sequence contains a PstI site at nucleotides 16220 through 16225. None of our other restriction enzymes have sites in the I27 sequence.
Carrion-Vazquez et al. list two vectors in their procedure, but I'm not sure about their respective roles.
pQE30
pQE30 (sequence) is listed as the "expression vector", but I'm not sure why they would need a non-expression vector, as they don't reference cross-vector subcloning after inserting their I27 monomer into the plasmid.
From the Qiagen site, the section around the linker nucleotides 115 through 203 is:
,RGS-His epitope__________________. ,BamHI.
Met Arg Gly Ser His His His His His His Gly Ser Ala Cys Glu Leu
ATG AGA GGA TCG CAT CAC CAT CAC CAT CAC GGA TCC GCA TGC GAG CTC
CGT CTC TTC GAT ACG ACA ACG ACA ACG ACA TTC GAA TAC GTA TCT AGA
,SmaI__.
,KpnI_. HindIII
Gly Thr Pro Gly Arg Pro Ala Ala Lys Leu Asn STOP
GGT ACC CCG GGT CGA CCT GCA GCC AAG CTT AAT TAG CTG AG
TTG CAA AAT TTG ATC AAG TAC TAA CCT AGG CCG GCT AGT CT
However, there is no BglII site in this linker. In fact, there is no BglII site in the entire pQE30 plasmid, so they'd need to use a third restiction enzyme to insert their I27 (which does contain a trailing BglII).
pUC19
From BCCM/LMBP and GenBank, the section around the linker nucleotides 233 through 289 is:
,SmaI_.
HindIII. ,PstI__. ,BamHI_. ,KpnI__.
Met STOP
AA GCT TGC ATG CCT GCA GGT CGA CTC TAG AGG ATC CCC GGG TAC CGA
GCT CGA ATT C
However, there is no BglII the entire pUC19 plasmid either, so they'd need to use a third restiction enzyme to insert their I27.
Questions
- Why do Carrion-Vazquez et al. list two different plasmids?
- What is the 3'-side restiction enzyme that Carrion-Vazquez et al. use to insert their I27 into their plasmid?
- What is the remote restriction enzyme that Carrion-Vazquez et al. use to break their opened plasmids (Kempe PstI equivalent).
- The BamHI and SmaI sites in pUC19 overlap, so it is unclear how you could use both to "linearize" pUC19. It would seem that either one would open the plasmid on its own, although I'm not sure you could "heal" the blunt-ended SmaI cut.
Since the Arg-Ser joint is formed by a BglII/BamHI overlap, why are there no BglII-coded amino acids after the last I27 in the I27RS₈ sequence? If there is, why do Carrion-Vazquez et al. not acknowledge it when they write [3]:
The full-length construct, I27RS₈, results in the following amino acid additions: (i) the amino-terminal sequence is Met-Arg-Gly-Ser-(His)6-Gly-Ser-I27 codons; (ii) the junction between the domains (BamHI-BglII hybrid site) is Arg-Ser; and (iii) the protein terminates in Cys-Cys.
Since they don't acknowledge an I27-Arg-Ser-Cys-Cys ending, might there be more amino acids in the C terminal addition?
Working backward
Since I'm stuck trying to get I27 into either plasmid, let's try and work backward from
Met-Arg-Gly-Ser-(His)₆-Gly-Ser-(I27-Arg-Ser)₇-I27-...-Cys-Cys
BglII/BamHI joint
The BglII/BamHI overlap would produce the expected Arg-Ser joint.
BglII BamHI
A + GATCC = AGATCC = Arg-Ser
TCTAG G TCTAGG
Final plasmid (pI27-8)
The beginning of this sequence looks like the start of pQE30's linker, so we'll assume the final plasmid was:
remote ... ,RGS-His epitope__________________. ,BamHI. I27...
... Met Arg Gly Ser His His His His His His Gly Ser Leu Ile ...
??? ... ATG AGA GGA TCG CAT CAC CAT CAC CAT CAC GGA TCC CTA ATA ...
??? ... CGT CTC TTC GAT ACG ACA ACG ACA ACG ACA TTC GAA GAT TAT ...
........I27 joint_. I27 ... final I27 ,BglII. continuation of pQE30?
... Glu Leu Leu ... Leu Arg Ser Cys Cys STOPSTOP...
... GAA TTG AGA TCC CTA ... TTG AGA TCT TGC TGC TAG TAG ...
... CTT AAC TCT AGG GAT ... GAT CTC GAG GTA GTA GCT GCT ...
Penultimate plasmid (pI27-4)
remote ... ,RGS-His epitope__________________. ,BamHI. I27...
Met Arg Gly Ser His His His His His His Gly Ser Leu Ile ...
??? ... ATG AGA GGA TCG CAT CAC CAT CAC CAT CAC GGA TCC CTA ATA ...
??? ... CGT CTC TTC GAT ACG ACA ACG ACA ACG ACA TTC GAA GAT TAT ...
... I27 joint_. I27 ... fourth I27 ,BglII. continuation of pQE30?
... Glu Leu Leu ... Leu Arg Ser Cys Cys STOPSTOP...
... GAA TTG AGA TCC CTA ... TTG AGA TCT TGC TGC TAG TAG ...
... CTT AAC TCT AGG GAT ... GAT CTC GAG GTA GTA GCT GCT ...
pI27-4 + BamHI + remote
remote ,BamHI. I27...
Leu Ile ...
? GA TCC CTA ATA ...
?? A GAT TAT ...
....... I27 joint_. I27 ... fourth I27 ,BglII. continuation of pQE30?
... Glu Leu Leu ... Leu Arg Ser Cys Cys STOPSTOP...
... GAA TTG AGA TCC CTA ... TTG AGA TCT TGC TGC TAG TAG ...
... CTT AAC TCT AGG GAT ... GAT CTC GAG GTA GTA GCT GCT ...
pI27-4 + BglII + remote
remote ... ,RGS-His epitope__________________. ,BamHI. I27...
Met Arg Gly Ser His His His His His His Gly Ser Leu Ile ...
?? ... ATG AGA GGA TCG CAT CAC CAT CAC CAT CAC GGA TCC CTA ATA ...
? ... CGT CTC TTC GAT ACG ACA ACG ACA ACG ACA TTC GAA GAT TAT ...
....... I27 joint_. I27 ... fourth I27 ,BglII.
... Glu Leu Leu ... Leu
... GAA TTG AGA TCC CTA ... TTG A
... CTT AAC TCT AGG GAT ... GAT CTC GA
pI27-8
remote ... ,RGS-His epitope__________________. ,BamHI. I27...
Met Arg Gly Ser His His His His His His Gly Ser Leu Ile ...
??? ... ATG AGA GGA TCG CAT CAC CAT CAC CAT CAC GGA TCC CTA ATA ...
??? ... CGT CTC TTC GAT ACG ACA ACG ACA ACG ACA TTC GAA GAT TAT ...
....... I27 joint_. I27 ... fourth I27 ,other. I27...
... Glu Leu Leu ... Leu Gly Ser Leu Ile ...
... GAA TTG AGA TCC CTA ... TTG AGA TCC CTA ATA ...
... CTT AAC TCT AGG GAT ... GAT CTC GAA GAT TAT ...
....... I27 joint_. I27 ... fourth I27 ,BglII. continuation of pQE30?
... Glu Leu Leu ... Leu Arg Ser Cys Cys STOPSTOP...
... GAA TTG AGA TCC CTA ... TTG AGA TCT TGC TGC TAG TAG ...
... CTT AAC TCT AGG GAT ... GAT CTC GAG GTA GTA GCT GCT ...
Continuing to the first plasmid, pI27-1 must have been
remote ... ,RGS-His epitope__________________. ,BamHI. I27...
... Met Arg Gly Ser His His His His His His Gly Ser Leu Ile ...
??? ... ATG AGA GGA TCG CAT CAC CAT CAC CAT CAC GGA TCC CTA ATA ...
??? ... CGT CTC TTC GAT ACG ACA ACG ACA ACG ACA TTC GAA GAT TAT ...
........I27 ,BglII. continuation of pQE30?
... Glu Leu Arg Ser Cys Cys STOPSTOP...
... GAA TTG AGA TCT TGC TGC TAG TAG ...
... CTT AAC CTC GAG GTA GTA GCT GCT ...
Potential pQE30 insertion points
- Kpn1 (present after BamHI in both plasmids)
Potential remote restriction enzymes
- BglI (pQE30 nucleotides 2583-2593 (GCCGGAAGGGC), Amp-resistance 3256-2396; pUC19 has two BglI sites (bad idea))
Available in a git repository.
Repository: pypid
Author: W. Trevor King
I've just finished rewriting my PID temperature control package in pure-Python, and it's now clean enough to go up on PyPI. Features:
- Backend-agnostic architecture. I've written a first-order process with dead time (FOPDT) test backend and a pymodbus-based backend for our Melcor MTCA controller, but it should be easy to plug in your own custom backend.
- The general PID controller will automatically tune your backend using any of a variety of tuning rules.
The README is posted on the PyPI page.
There are a number of open source packages dealing with aspects of single-molecule force spectroscopy. Here's a list of everything I've heard about to date.
| Package | License | Purpose |
|---|---|---|
| calibcant | GPL v3+ | Cantilever thermal calibration |
| fs_kit | GPL v2+ | Force spectra analysis pattern recognition |
| Hooke | LGPL v3+ | Force spectra analysis and unfolding force extraction |
| sawsim | GPL v3+ | Monte Carlo unfolding/refolding simulation and fitting |
| refolding | Apache v2.0 | Double-pulse experiment control and analysis |
calibcant
Calibcant is my Python module for AFM cantilever calibration via the thermal tune method. It's based on Comedi, so it needs work if you want to use it on a non-Linux system. If you're running a Linux kernel, it should be pretty easy to get it running on your system. Email me if there's any way I can help set it up for your lab.
fs_kit
fs_kit is a package for force spectra analysis pattern
recognition. It was developed by Michael Kuhn and Maurice Hubain at
Daniel Müller's lab when they were at TU Dresden
(paper). It has an Igor interface, but the bulk
of the project is in C++ with a wxWidgets interface. fs_kit
is versioned in CVS at bioinformatics.org, and you can check out
their code with:
$ cvs -d:pserver:anonymous@bioinformatics.org:/cvsroot checkout fskit
The last commit was on 2005/05/16, so it's a bit crusty. I patched things up back in 2008 so it would compile again,
0001-Added-math.h-include-to-fs align histogram2d.h.patch
Posted Sat Apr 23 15:00:17 2011
0002-changed-abs-double-to-fabs-double-in-fs fit spectr.patch
Posted Sat Apr 23 15:00:17 2011
0003-Updated-wxWindows-code-to-compile-on-wx-2.8.patch
Posted Sat Apr 23 15:00:17 2011
0004-Added-wxglade-entry-to-Makefile-for-regenerating-aut.patch
Posted Sat Apr 23 15:00:17 2011
but when I emailed Michael with the patches I got this:
On Thu, Oct 23, 2008 at 11:21:42PM +0200, Michael Kuhn wrote:
> Hi Trevor,
>
> I'm glad you could fix fs-kit, the project is otherwise pretty dead,
> as was the link. I found an old file which should be the tutorial,
> hopefully in the latest version. The PDF is probably lost.
>
> bw, Michael
So, it's a bit of a fixer-upper, but it was the first open source package in this field that I know of. I've put up a PDF version of the tutorial Michael sent me in case you're interested.
Hooke
Hooke is a force spectroscopy data analysis package written in Python. It was initially developed by Massimo Sandal, Fabrizio Benedetti, Marco Brucale, Alberto Gomez-Casado while at Bruno Samorì's lab at U Bologna (paper; surprisingly, there are commits by all of the authors except Samorì himself). Hooke provides the interface between your raw data and theory. It has a drivers for reading most force spectroscopy file formats, and a large number of commands for manipulating and analyzing the data.
I liked Hooke so much I threw out my already-written package that had been performing a similar role and proceeded to work over Hooke to merge together the diverging command-line and GUI forks. Unfortunately, my fork has not yet been merged back in as the main branch, but I'm optimistic that it will eventually. The homepage for my branch is here.
sawsim
While programs like Hooke can extract unfolding forces from velocity-clamp experiments, the unfolding force histograms are generally compared to simulated data to estimate the underlying kinetic parameters. Sawsim is my package for performing such simulations and fitting them to the experimental histograms (paper). The single-pull simulator is written in C, and there is a nice Python wrapper that manages the thousands of simulated pulls needed to explore the possible model parameter space. The whole package ends up being pretty fast, flexible, and convenient.
refolding
Refolding is a suite for performing and analyzing
double-pulse refolding experiments. It was initially developed by
Daniel Aioanei, also at the Samorí lab in Bologna (these guys are
great!). The experiment-driver is mostly written in Java with the
analysis code in Python. The driver is curious; it uses the
NanoScope scripting interface to drive the experiment through the
NanoScope software by impersonating a mouse-wielding user (like
Selenium does for web browsers). See the RobotNanoDriver.java
code for details.
My wife was recently reviewing some pulse oxymeter notes while working a round of anesthesia. It took us a while to trace the logic through all the symbol changes and notation shifts, and my wife's math-discomfort and my bio-discomfort made us more cautious than it turns out we needed to be. There are a number of nice review articles out there that I turned up while looking for explicit derivations, but by the time I'd found them, my working notes had gotten fairly well polished themselves. So here's my contribution to the pulse-ox noise ;). Highlights include:
- Short and sweet (with pictures)
- Symbols table
- Baby steps during math manipulation
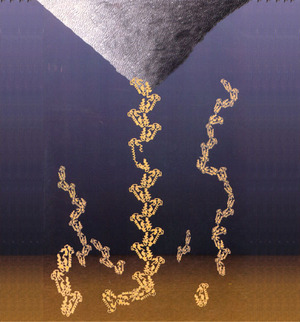
Force spectroscopy is the process of extracting information about the unfolding (or unbinding) characteristics of a protein (or ligand-receptor pair) by measuring force vs. extension curves while gradually ripping the protein (or pair) apart. Consider this cartoon representation of the procedure
The AFM tip is pulling a protein chain away from the substrate, causing one of the protein domains to uncoil.
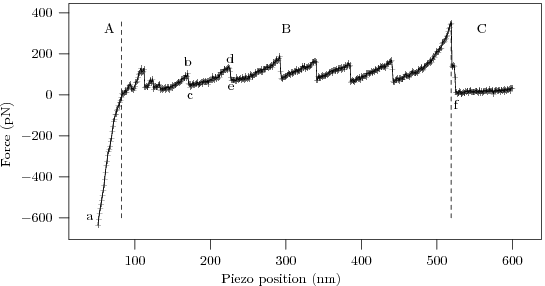
The procedure yields 'force curves' like this
To interpret the force curve, let us examine it piece-by-piece as the AFM tip gradually pulls away from the substrate.
- The linear 'contact' region demonstrates the Hooke's law behavior of the AFM cantilever, with force ∝ displacement.
- The high force 'bulge' linking the contact region to the sawtooth comes from the AFM tip pulling free of the surface and associated protein 'mat' (the cartoon being excessively pretty, and our sample having too high a protein concentration :p).
- The characteristic 'sawtooth' comes from several identical domains unfolding one after the other.
- After the last of the protein domains unfolds the protein snaps off of the AFM tip (or the substrate), and the deflection of the now-free cantilever ceases to depend on distance.
The clickloc.tk micro-app just opens an image file and prints out the pixel coordinates of any mouse clicks upon it. I use it to get rough numbers from journal figures. You can use scale click.sh to convert the output to units of your choice, if you use your first four clicks to mark out the coordinate system (xmin,anything), (xmax,anything), (anything,ymin), and (anything,ymax).
$ pdfimages article.pdf fig
$ clickloc.tk fig-000.ppm > fig-000.pixels
$ scale_click.sh 0 10 5 20 fig-000.pixels > fig-000.data
Take a look at plotpick for grabbing points from raw datafiles (which is more accurate and easier than reverse engineering images).
Available in a hg repository.
Repository: hooke
Author: W. Trevor King
Hooke is a force spectroscopy data analysis package. For example, Hooke can extract unfolding forces from your experimental data. You can then fit the unfolding forces to models using my sawsim simulator. Of course, some experiments (e.g. force clamp) need no Monte Carlo analysis, so for those, Hooke alone provides a complete analysis package.
Getting started
I've tested Hooke on Gentoo and Debian, and I've got an ebuild in my Gentoo overlay. It should also run fine on Windows, etc., but I don't have easy access to Windows boxes with Python, so I don't test it there as often.
Available in a git repository.
Repository: sawsim
Author: W. Trevor King
Introduction
My thesis project investigates protein unfolding via the experimental technique of force spectroscopy. In force spectroscopy, we mechanically stretch chains of proteins, usually by pulling one end of the chain away from a surface with an AFM.
For velocity clamp experiments (the simplest to carry out experimentally), the experiments produce "sawtooth" force-displacement curves. As the protein stretches, the tension increases. At some point, a protein domain unfolds, increasing the total length of the chain and relaxing the tension. As we continue to stretch the protein, we see a series of unfolding peaks. The GPLed program Hooke analyzes the sawtooth curves and extracts lists of unfolding forces.
Lists of unfolding forces are not particularly interesting by themselves. The most common approach for extracting some physical insights from the unfolding curves is to take a guess at an explanatory model and check the predicted behavior of the model against the measured behavior of the protein. If the model does a good job of explaining the protein behavior, it might be what's actually going on behind the scenes. Sawsim is my (published!) tool for simulating force spectroscopy experiments and matching the simulations to experimental results.
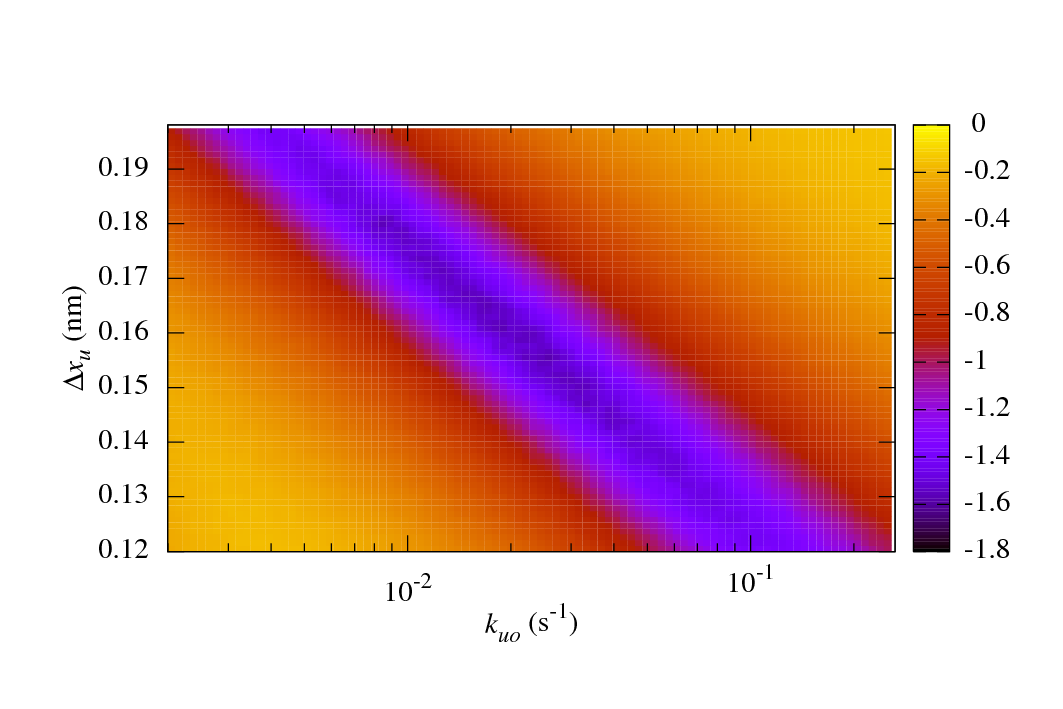
The main benefits of sawsim are its ability to simulate systems with arbitrary numbers of states (see the manual) and to easily compare the simulated data with experimental values. The following figure shows a long valley of reasonable fits to some ubiquitin unfolding data. See the IJBM paper (linked above) for more details.
Getting started
Sawsim should run anywhere you have a C compiler and Python 2.5+. I've tested it on Gentoo and Debian, and I've got an ebuild in my Gentoo overlay. It should also run fine on Windows, etc., but I don't have access to any Windows boxes with a C compiler, so I haven't tested that (email me if you have access to such a machine and want to try installing Sawsim).
Astrometry.net is awesome.