A Simplified Representation of
Anisotropic Charge Distributions
Drexel Physics Colloquium
Travis Hoppe , Allen Minton
NIH, NIDDK, LBG
Main Idea
Capture structural features of protein solutions.
Phase Changes &
Virial Coefficients
via
pH Dependence
Concentration Dependence
Prior Work
Coarse-grained models:
Hard spheres, point charges/dipoles, LJ liquids
Fine-grained models:
All-atom molecular dynamics, quantum ab-initio
Liquid theory
Ornstein-Zernike equation, Percus-Yevick closures...
Anisotropy
Directional (angular) dependence
eg. polarizations, nematic phases, chirality, ...
The Process
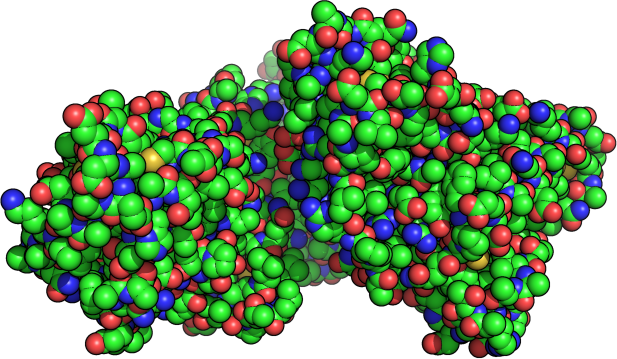
The Process
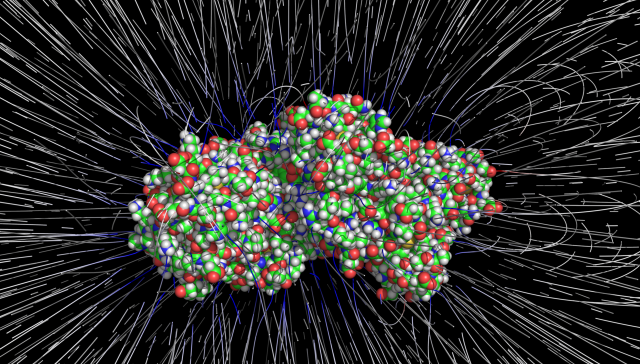
The Process
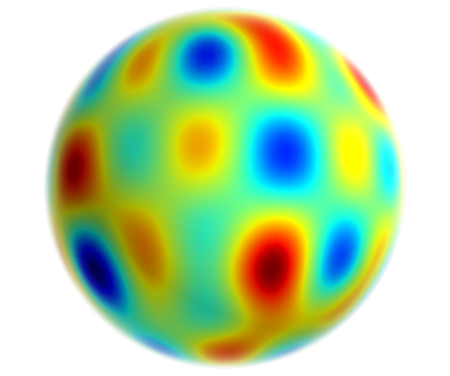
Spherical Harmonic decomposition
The Process
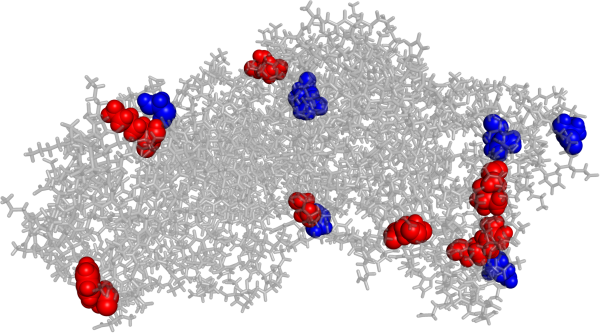
The Process
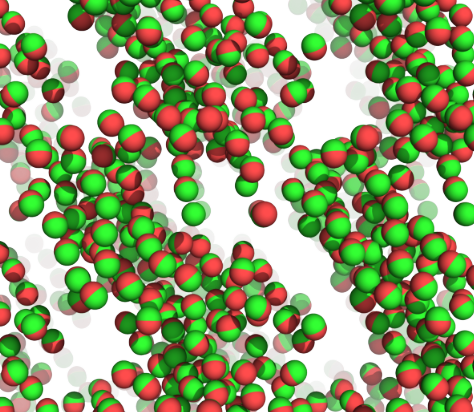

Electrostatic field
Coulomb's Law (point charge)
Correction for dielectrics?
What to do with the solvent?
Yukawa Potential
First order approximation to screening effects.
Charge strength decays exponentially due to ions.
Poisson Boltzmann
Describes the electrostatic interaction between a charge distribution and an ionic solution.
Assumes ions are Boltzmann-distributed in the solution.
Can be linearized and solved on a computer efficiently.
Splits space into regions of discrete  .
.
Adaptive Poisson-Boltzmann Solver
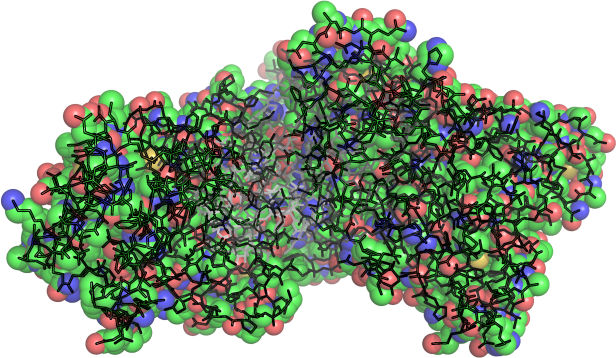


Simplify!
Build a basis set
Represent the data with a set of operators with the same boundary conditions. In effect, reduce millions of data points to dozens .
1D: Harmonics
1D: Bessel Functions
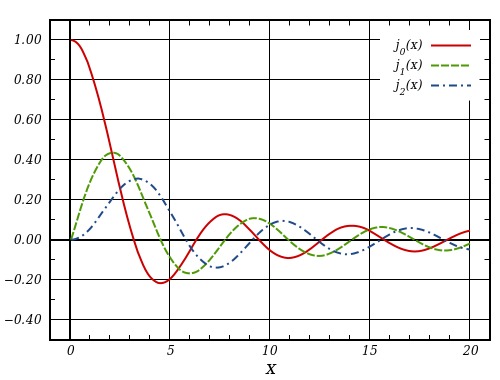
3D: Spherical Harmonics

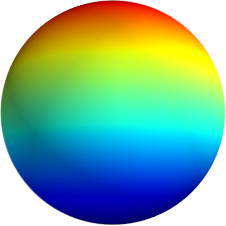
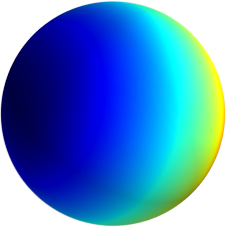
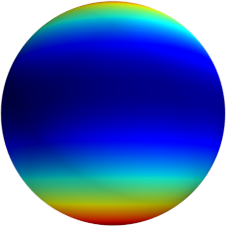
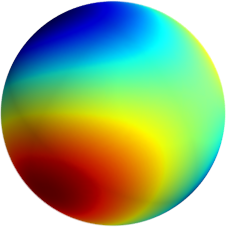
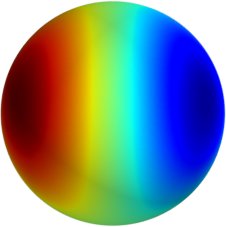
3D: Spherical Harmonics
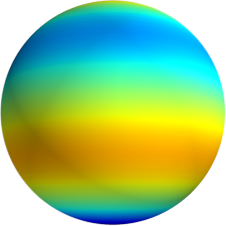
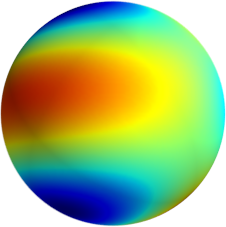
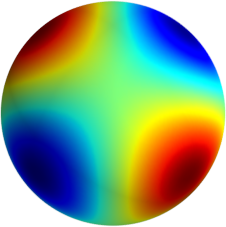
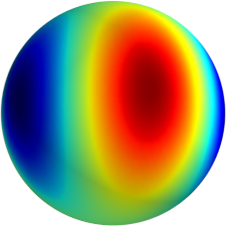
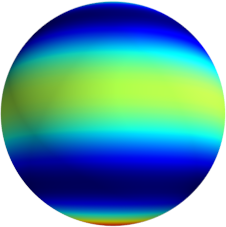
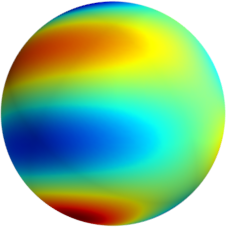
3D: Spherical Harmonics
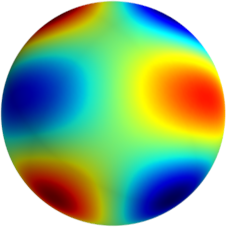
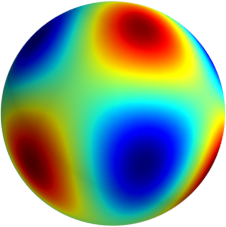
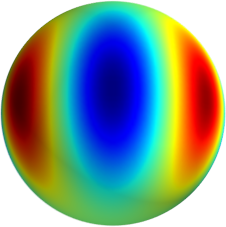
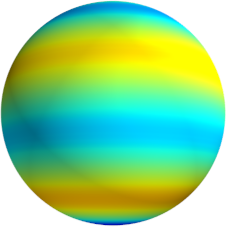
Approximate  as a function of the spherical harmonics...
as a function of the spherical harmonics...
 +
+  +
+  +
+  +
+  + ...
+ ...





Accuracy Test
Define an error between the target potential 
and the generated potential 
Does the Spherical Harmonics work?
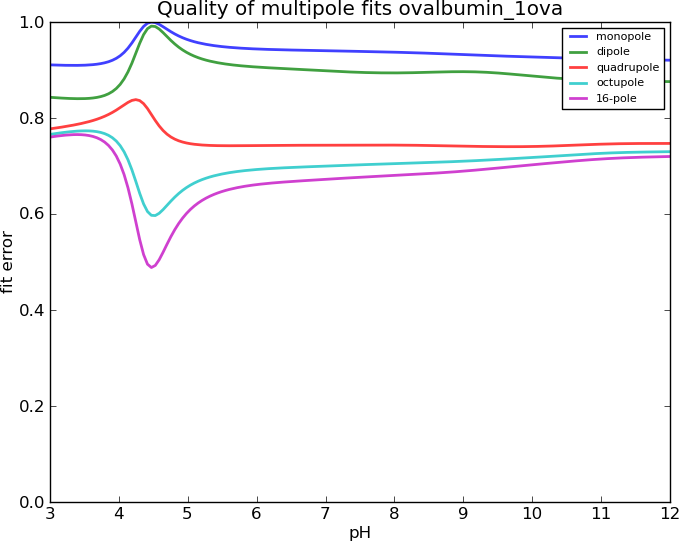
Why do the Spherical Harmonics fail?
They are expansions of a field 
When we consider the exponential decay 
We need to use Bessel functions
Ovalbumin, I=0.15M
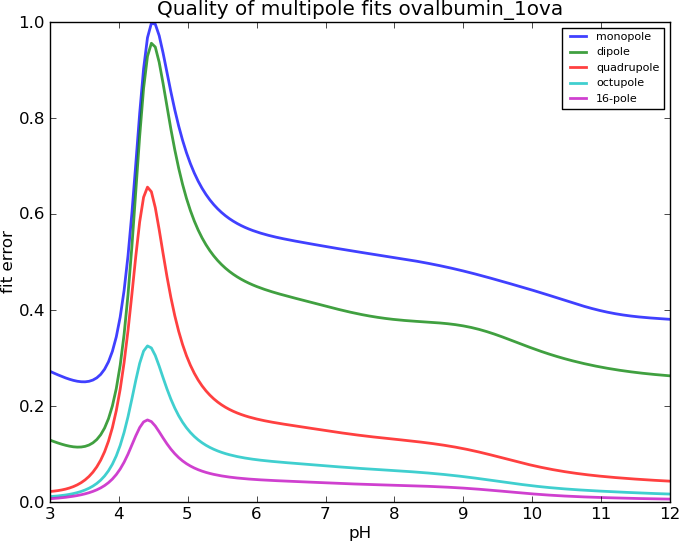
Lysozyme, I=0.15M
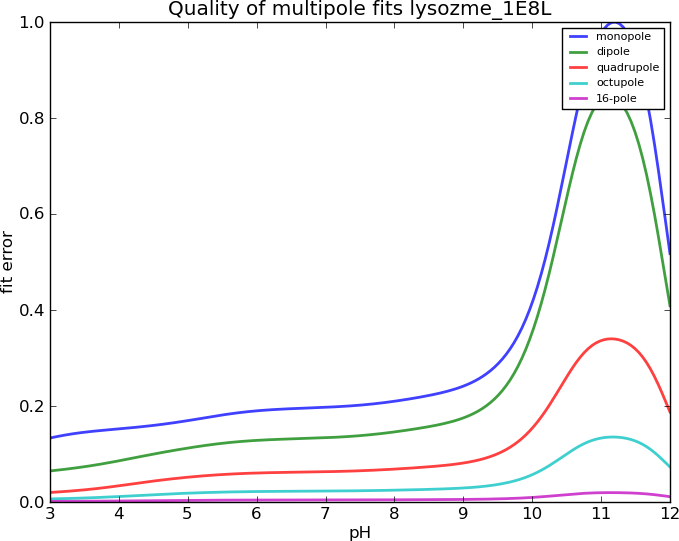
Human Serum Albumin, I=0.15M
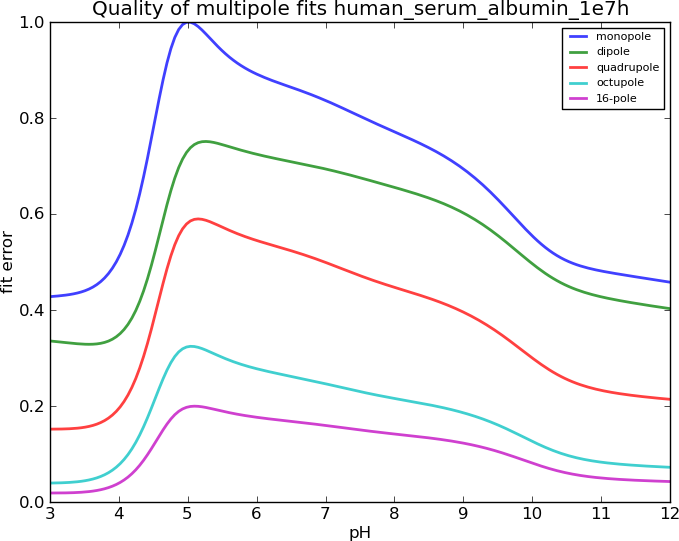
Ovalbumin, I=0.30M
Results degrade with increasing ionic strength
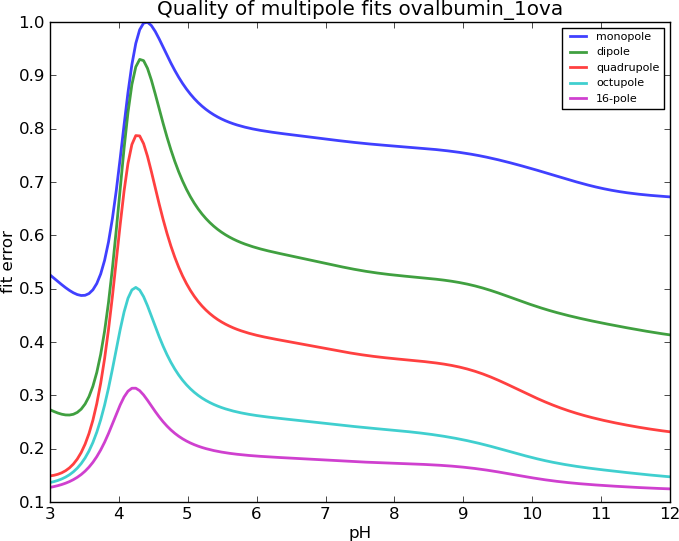
Ovalbumin, I=0.45M
Results degrade with increasing ionic strength
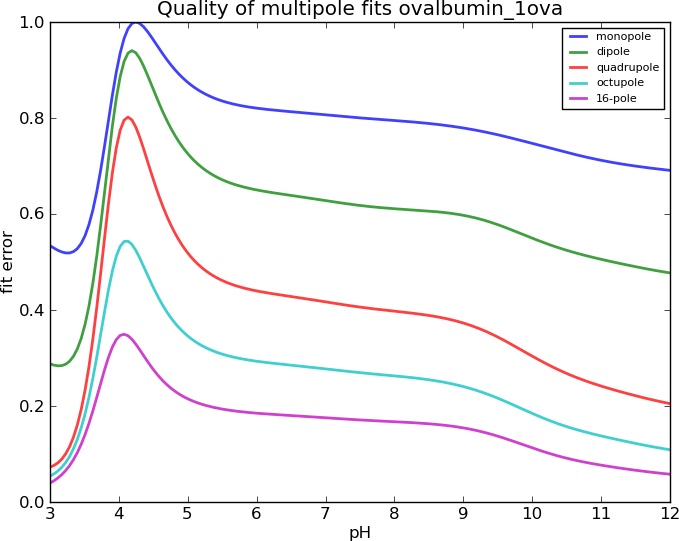
 , breaks down
, breaks down
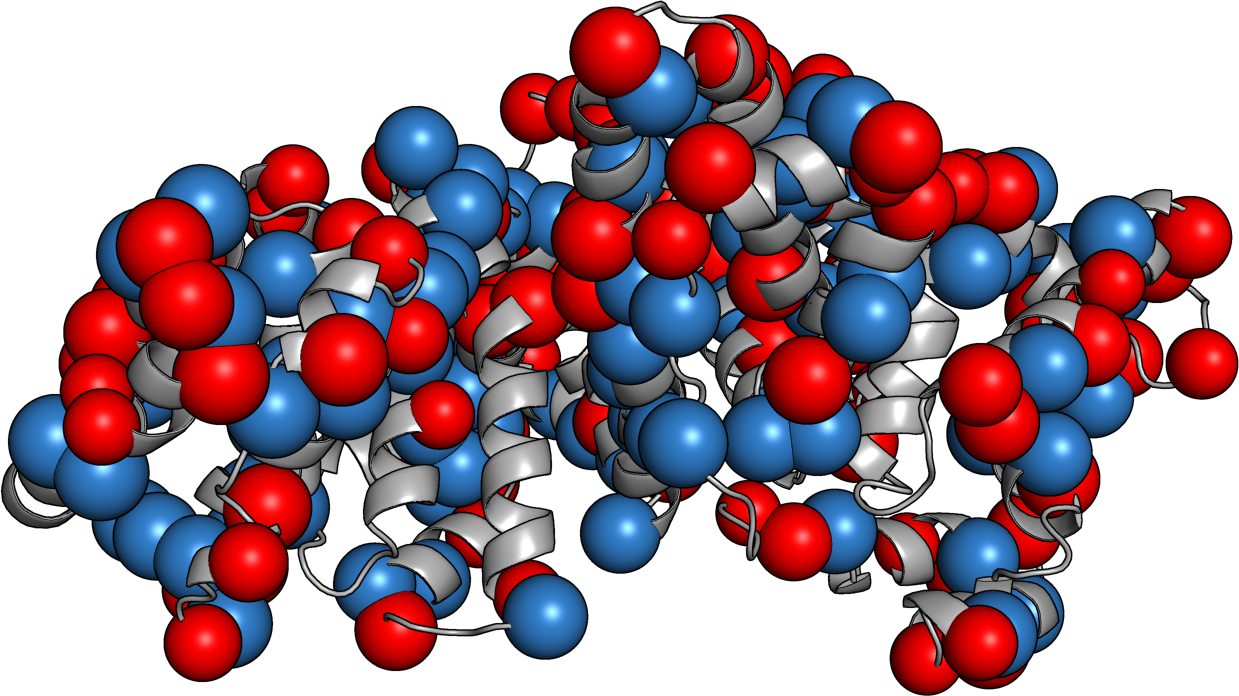
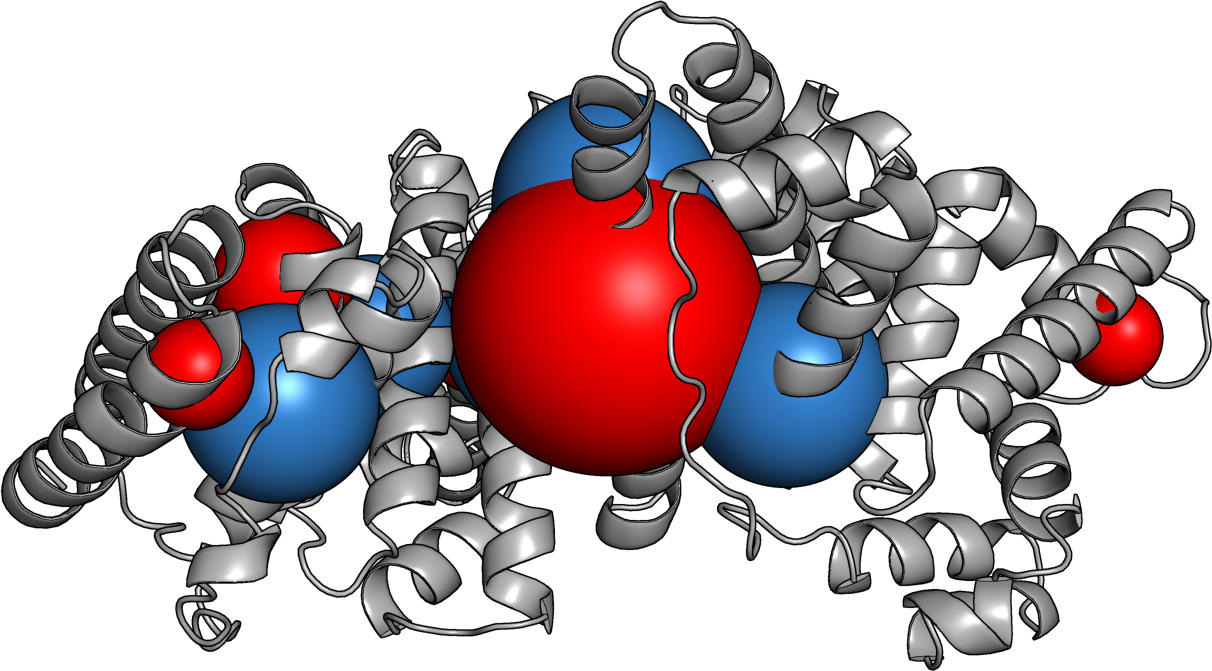
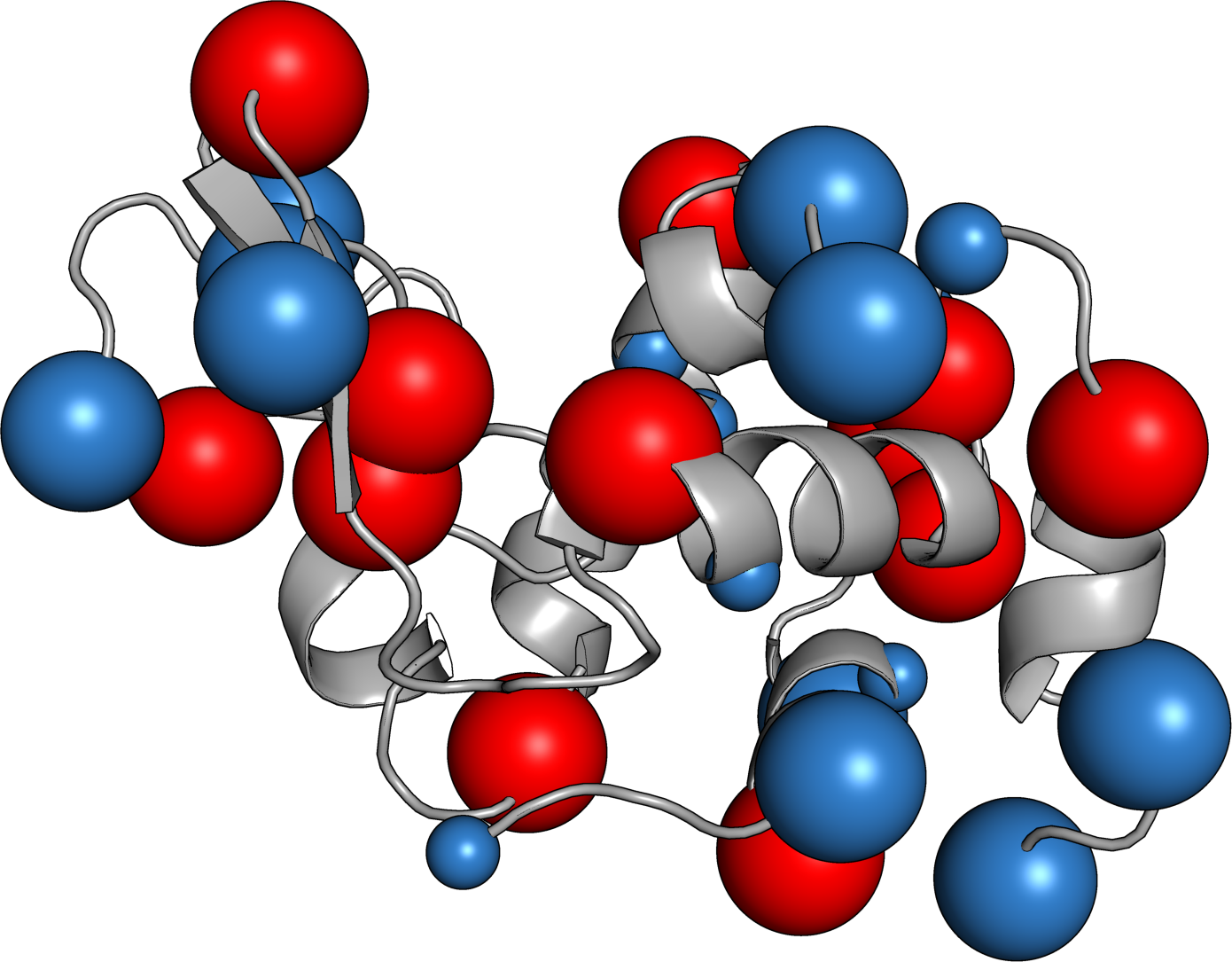
Protein Caricatures
Good approximation of the near field, poor up close
Captures the anisotropic field
especially near the isoelectric point
Macrocharge approximations make for reasonable
models of large protein solutions
Charge fitting
Fit discrete macrocharges to match harmonics
Potential is linear in charge magnitude -
only coordinates need to be fit
Why not use multipoles directly?
Extremely complicated for anything > quadrupole
Macrocharges mapped to protein coordinates,
useful in it's own right?
Fitting Problems
Charges must be constrained
Outside the expansion, the fits are not valid
Creation of artificial dipoles
Unstable w.r.t. fits, constant  results in extreme magnitude fluctuations
results in extreme magnitude fluctuations
Effective charge error, Lysozyme
 are unit normalized coefficients for APBS and effective charges
are unit normalized coefficients for APBS and effective charges
Effective charge error
How many charges needed?
Human Serum Albumin
Human Serum Albumin
Lysozyme
Lysozyme
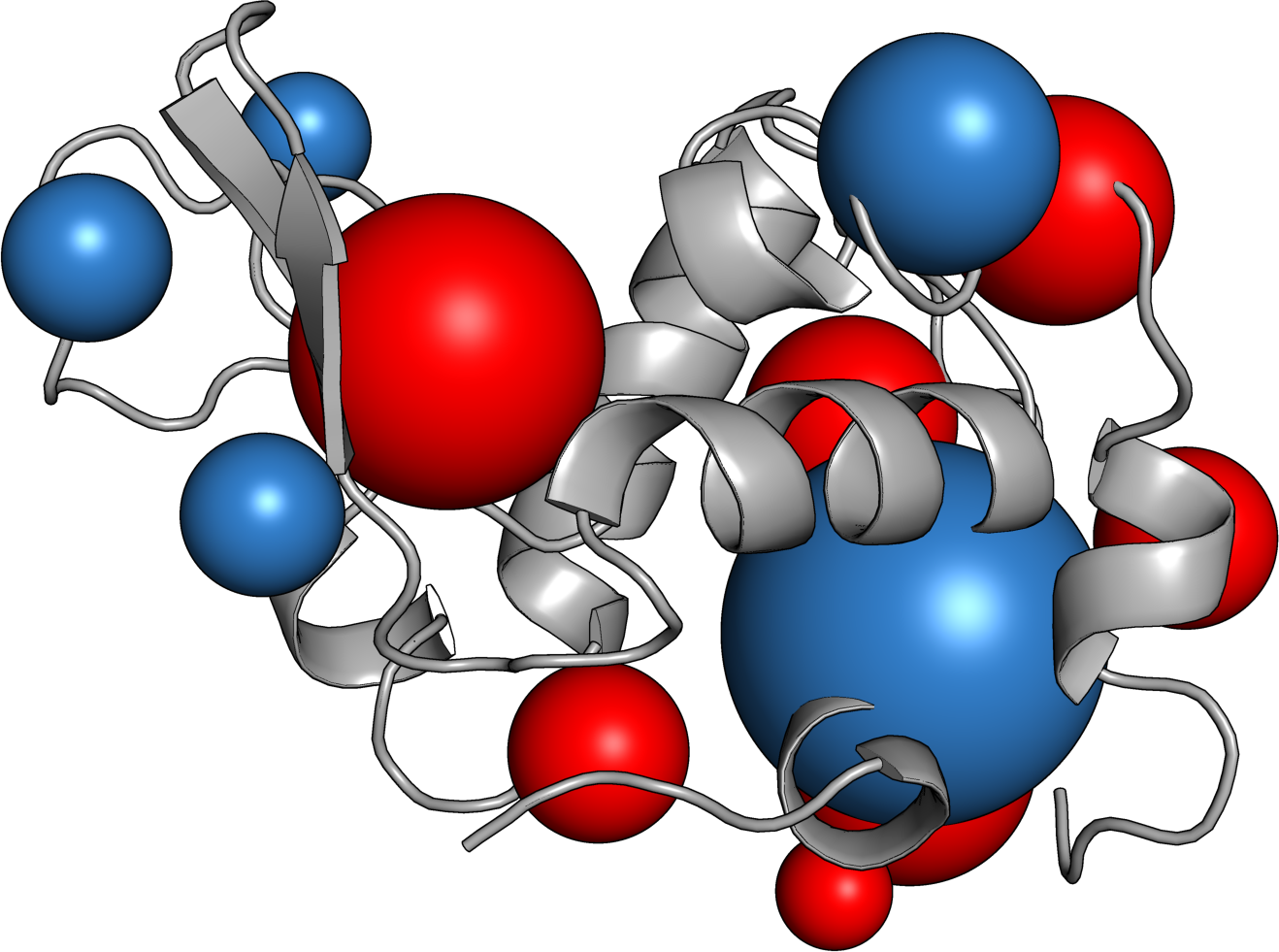
Ovalbumin
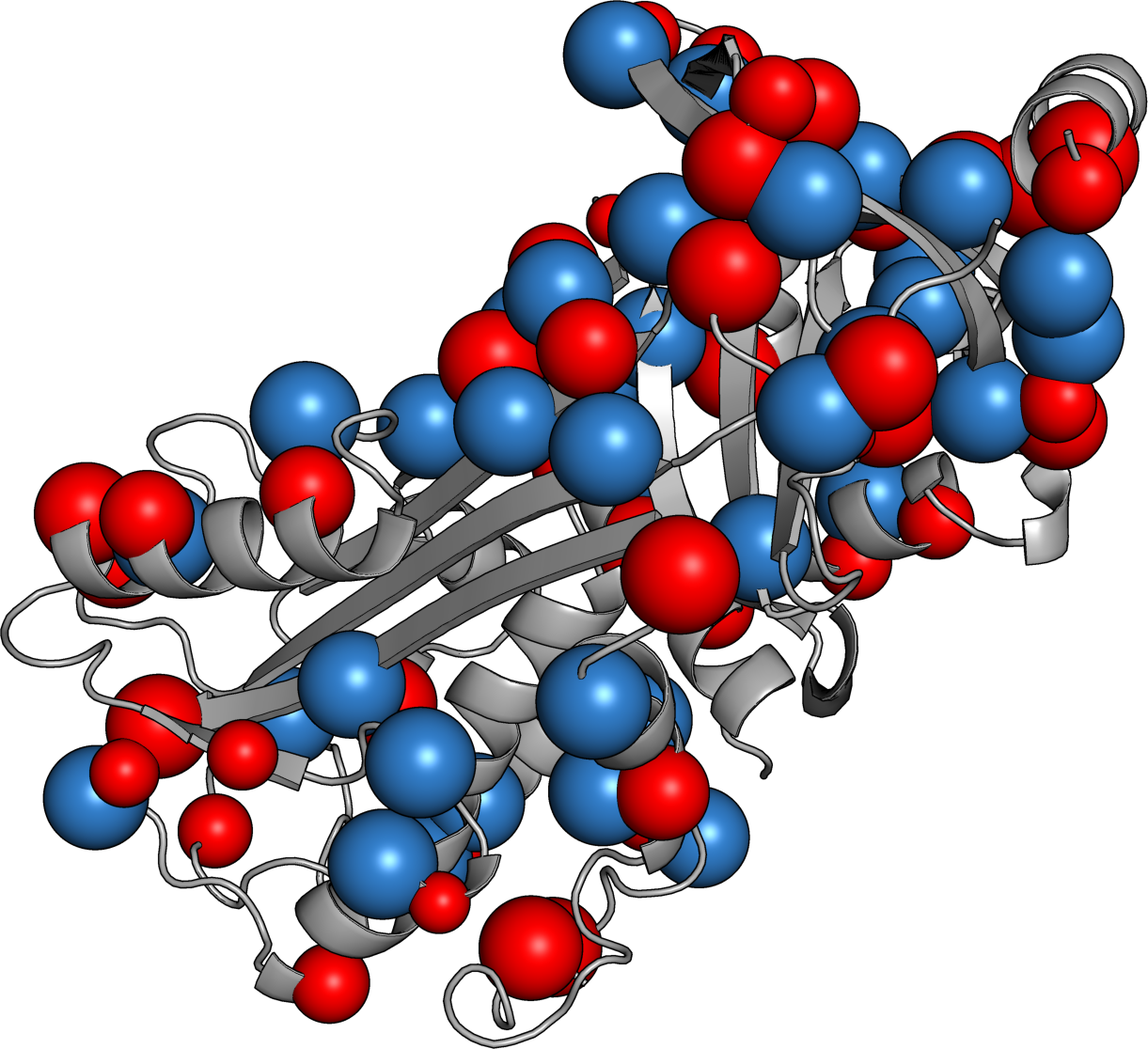
Ovalbumin
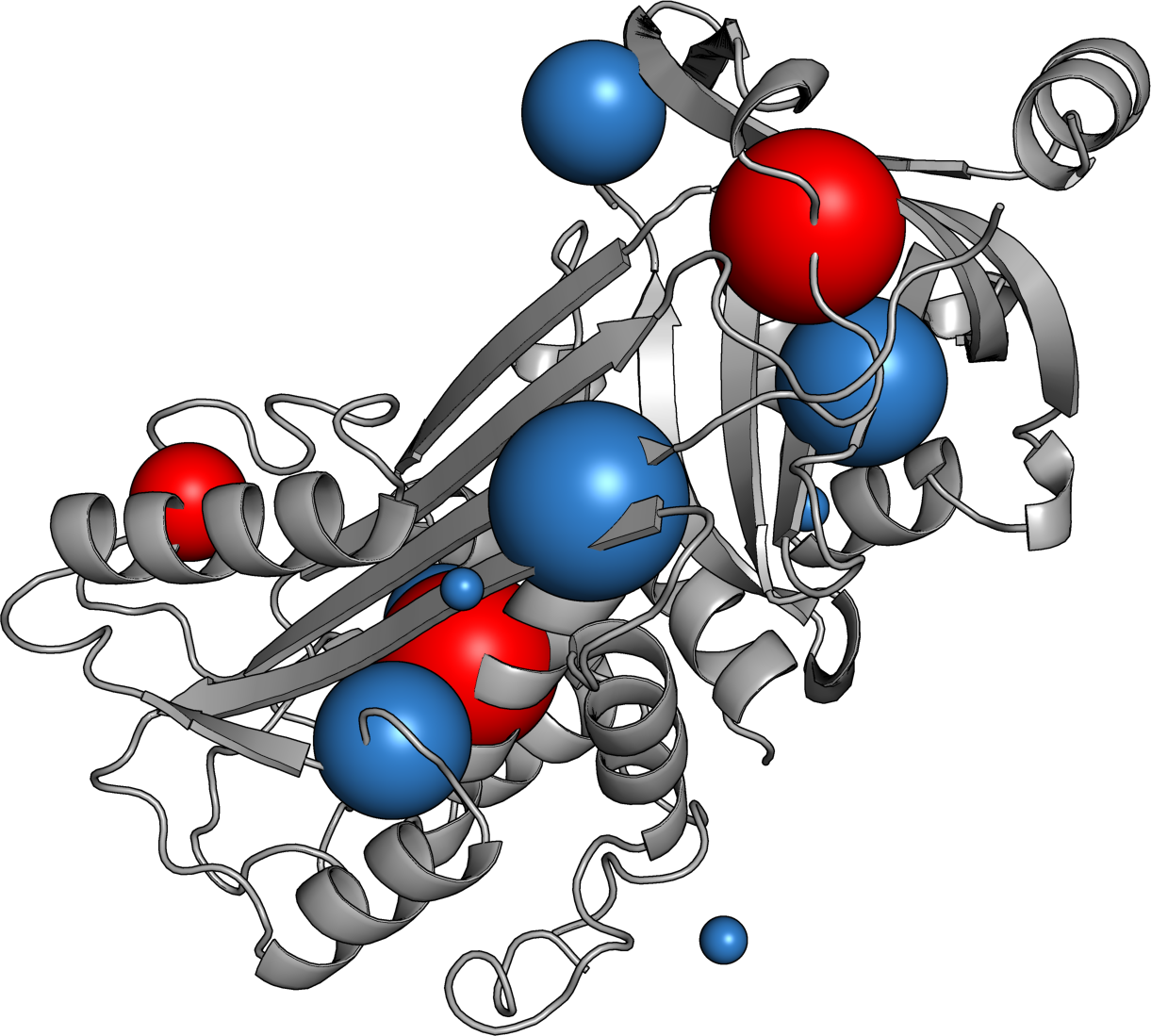
So Far...
Built a coarse-grained representation of a protein in an ionic solution for a given pH.
What's next?
Interaction Energy
Second Virial Coeff, 

Predictions
Phase separations

Phase separations lead to sudden fundamental changes in liquid structure and local density
This is usually really important
Predictions
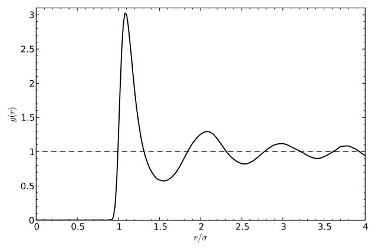
Radial distributions & B2
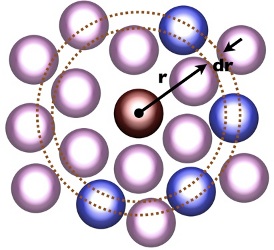
Basic Science
Match up experiential data with results of computational models
Classify common protein solution behavior from macrocharges or multipoles
Extrapolate to mutations and other unknown proteins
Thank you
How were these slides made?
Math Rendering: 
JavaScript: reveal.js
Custom Markdown: Travis Hoppe
What does Markdown look like?
A text-based human-readable markup.
Equation rendering is simple  .
.
The code for this particular slide looks like this:
## What does Markdown look like?A **text-based** human-readable markup.
Equation rendering is simple $e^{i \pi} = -1$.
The code for this *particular slide* looks like this:


![\vec{\nabla}\cdot\left[\epsilon(\vec{r})\vec{\nabla}\phi(\vec{r})\right] = -\rho^{f}(\vec{r}) - \sum_{i}c_{i}^{\infty}z_{i} q \lambda(\vec{r}) \exp \left[{\frac{-z_{i}q\phi(\vec{r})}{k_B T}}\right]](equations/baf54ae6b0a4ce36b56f4a0c3ef390ad8cbc371bff1373da1a242e11592d2a4a.png)


